Replication
Copying: Cool for DNA, Cool for Us
Back when some of Shmoop's classmates were in school, their teachers would always catch them copying. But, the classmates told their teachers that it was in their DNA to copy. She could not argue with that logic, so they never got into trouble. What a nice little story.What parallel are we trying to draw with this not-so-subtle and hokey tale? DNA copies all the time, and it must if it wants to keep making all those cells. And, oh, how it does! DNA replication is the copying of DNA so that replicating cells will have enough DNA for daughter cells, or the new cells derived from the original cell. Cell division, or the making of one cell into two new cells, occurs through mitosis for non-sex cells, or those cells not involved in...sex, and meiosis for sex cells, or those cells that do get down and dirty faster than Vinny and Pauly D on a good night at Karma.
It's Just a Phase: Mitosis or Meiosis
Though cells come in all shapes and sizes, like a Benetton ad, there are basically two types of cells:- Sex cells (no, not those things in Amsterdam)
- Somatic cells (non-sex cells, which are also not the things in Amsterdam)
- Sperm cells that contain the male half of DNA
- Egg cells that contain the female half of DNA
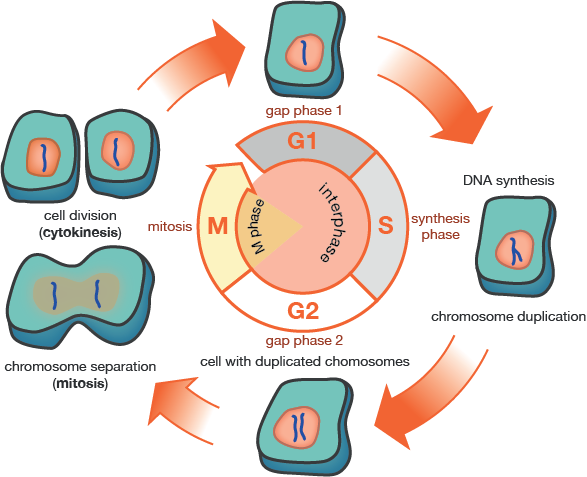
There are four major stages in the life cycle of most eukaryotic cells (excluding sex cells).
These are called
- Gap 1 (or G1) phase
- Synthesis (or "S") phase
- Gap 2 (G2) phase
- Mitosis
G1
G1 is the phase of the eukaryotic life cycle after mitosis and before DNA synthesis, which is S phase. During mitosis, most of the non-mitosis activity has been shut down, and G1 is the period where everything starts again. G1 is like a crazed insomniac that wakes the cell by saying, "Hey, I have been cutting out coupons all night, time to wake up and play with me!"S Phase
S phase is the DNA replication phase of the life cycle. We will talk more about this later, but if you cannot wait, click ahead. This phase is where the cell doubles the amount of DNA to make enough DNA for the daughter cells, and the cell goes from 2n to 4n, where n is the number of sets of chromosomes, or ploidy. Most somatic cells are 2n, or diploid, while sex cells are haploid, meaning that they have half as much DNA as somatic cells (regular cells).G2 Phase
G2 is the Jan Brady of the cell cycle. Everyone always goes on about "Mitosis, Mitosis, Mitosis!" and nobody cares about poor G2. It is the period of the cell cycle where more protein synthesis occurs and microtubules are made, which are important for mitosis. "That's it?" You ask? Yup. Not really that important, which is why many cells skip this step. Frog cells skip it, as well as many cancerous cells. Poor G2…Mitosis
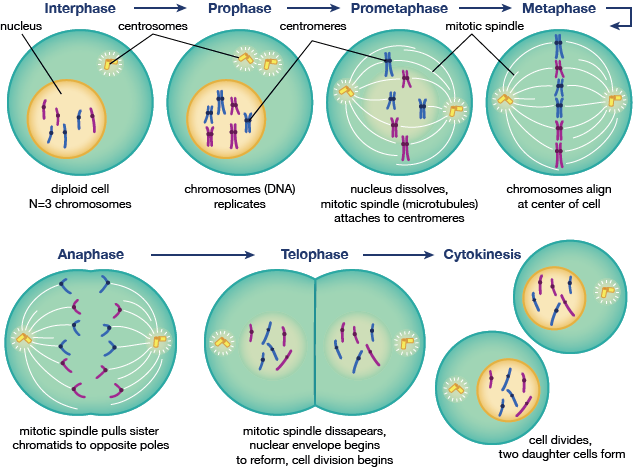
Mitosis is the stage in the cell cycle that is so important, it has a cycle of its own. Mitosis begins after G2, and starts with prophase, or the condensation of chromatin into chromosomes and centrosomes moving toward opposing poles of the cell. This step is followed by the prometaphase stage, an intermediate stage between prophase and metaphase, where the nuclear envelope breaks down and chromosomes start assembling onto microtubles connecting the centrosomes. Metaphase is where all the chromosomes are aligned in the center, which is followed by anaphase, where the chromosomes are split in half, and daughter chromatids move to the closest centrosome pole. Telophase and cytokinesis are the final steps of cell division and reassembly of the nucleus into two daughter cells, respectively.
Meoisis
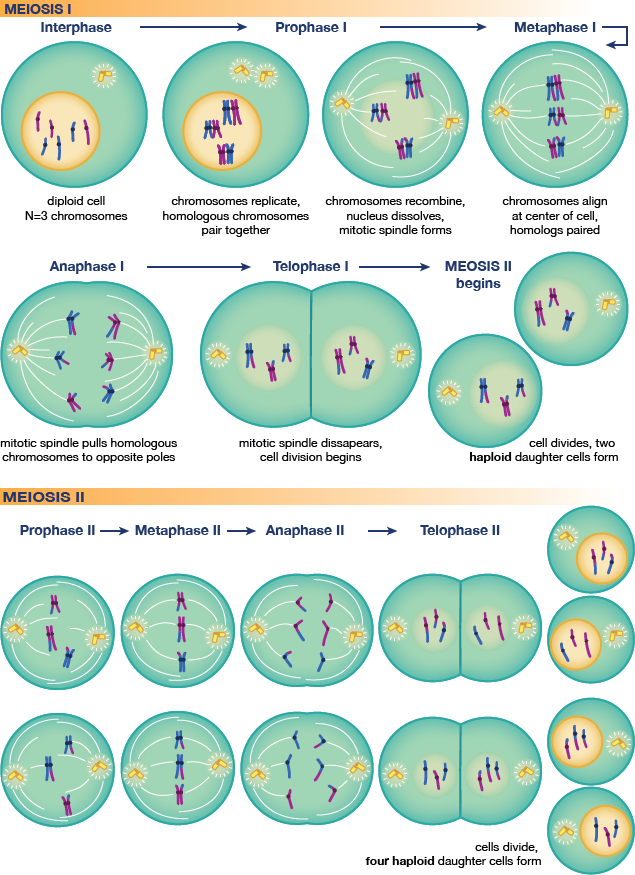
Meiosis is the M phase for sex cells. It begins like mitosis, and after the production of the first two daughter cells, it continues a second mitosis cycle. The 2n daughter cells divide further to produce 4 daughter cells, and a lot of overpriced weddings, each with n chromosomes.
In the Beginning…DNA Replication Origins
If DNA never replicated, meiosis and mitosis would slowly halve the size of the genome until each cell would die, which probably would not take long. Therefore, it is important that DNA doubles itself to account for the cells splitting during mitosis/meiosis. DNA replication is similar to RNA transcription.Below is a table comparing the two:
| DNA Replication | RNA Transcription | |
| Product | Double-stranded DNA | mRNA |
| Enzyme | DNA polymerase | RNA polymerase |
| Nucleotides Added | Deoxynucleotide triphosphates | Nucleotide triphosphates |
| Template Strand | Both | Both, but only transcribes from antisense strand |
Origins differ depending on the type of DNA. Most eukaryotic cells and some bacteria and viruses have linear DNA, meaning that there is a beginning and an end to the DNA sequence. Therefore, the origins of replication are the 5' end of the linear DNA. Most bacteria and some viruses have circular DNA, which means that there is a specific sequence that DNA replication proteins bind to (which is somewhat analogous to RNA transcription), and replication begins from this site.
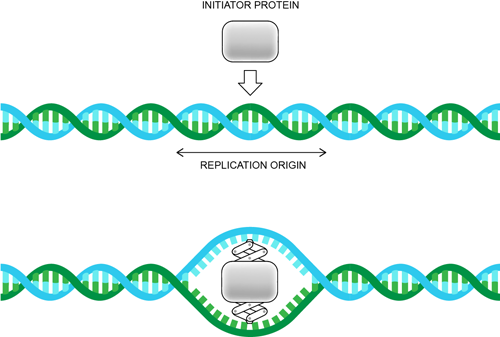
DNA replication starts with an initiator protein binding, such as dnaA in E. coli bacteria, or a complex of proteins, like the origin recognition complex in yeast. The function of initiators is to pry apart the two strands of DNA so that replication can begin. This situation is not unlike a jack lifting your car when you change a tire. (Because we all know how to do that, right?) If you don't know how to change a tire, then call a tow company and ask them how they do it. After they tell you how, scream: "That's exactly like DNA Replication!" hang up the phone, and then pray that you never need to change a tire again.
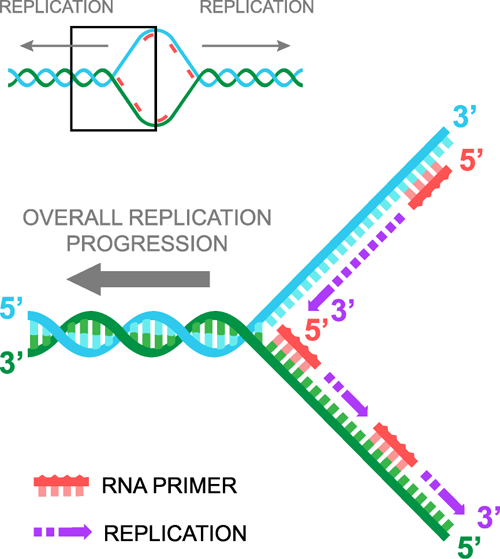
Once the DNA has been pried apart, RNA primers enter, one binds to each strand of DNA, and then DNA replication proceeds in a 5' to 3' direction. It is important to realize that there are two 5' to 3' replications happening simultaneously. These two replications are each called a replication fork. Not to be confused with the duplication spoon.

Pass the Replication Fork
When you split DNA, we have a strand that is 5' to 3' as we look left to right, which we will call the "lagging" strand and a complementary strand that reads 3' to 5' as we look left to right, which we will call the "leading" strand. Therefore, when we add RNA primers, one primer binds to the leading strand, and the other primer binds to the lagging strand. The primer that binds to the leading strand is identical to the sequence of the lagging strand, while the primer that binds to the lagging strand is identical to the leading strand sequence. We hope your head has not exploded yet.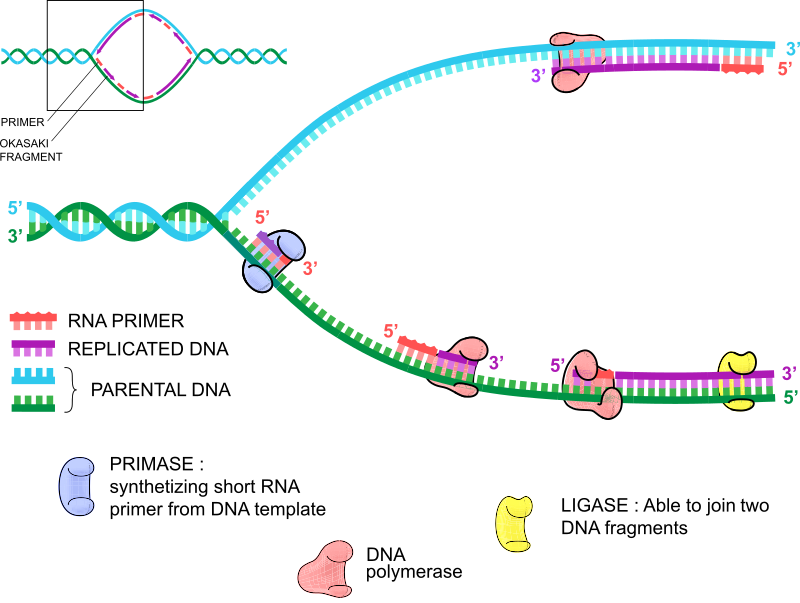
Think of DNA replication like your left and right hands. They are essentially the same but have subtle differences. Your left hand has the thumb as the fifth finger if you count left to right, while your right hand has the thumb as the first finger. See? Sorry if we blew your mind there. Therefore, like your hands, both strands of DNA are replicating in the same way, but there is a subtle difference. DNA replication only works in a 5' to 3' direction, so two DNA polymerase molecules bind the RNA primers and start replicating in opposite directions. The leading strand replication drives the replication process because the RNA primer drives replication of the 5' to 3' strand.
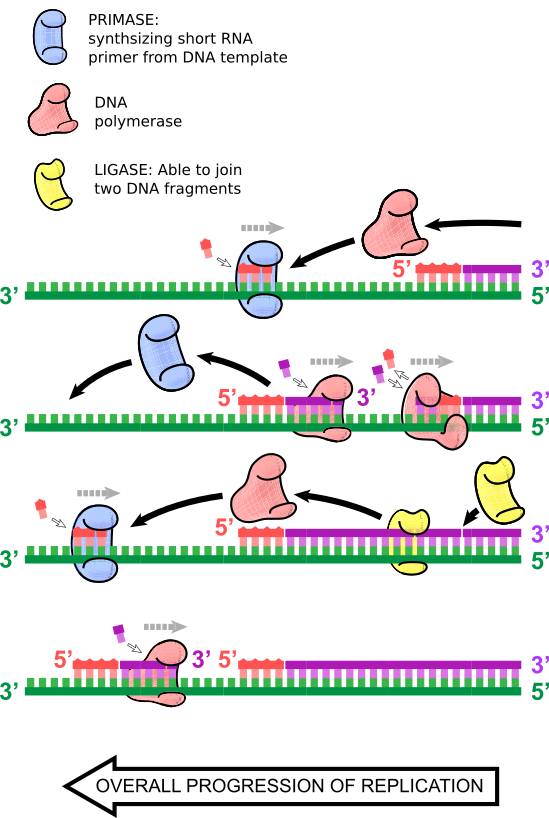
"But Shmoop, how do you replicate the 'lagging' strand?" you might ask.
As its name suggests, DNA replication of the leading strand continues for some time until a new RNA primer inserts, and a second DNA polymerase primes from that second lagging RNA primer and replicates DNA until it hits the first RNA primer, or previous stretch of double-stranded DNA. There, DNA polymerase falls off, and DNA ligase joins these fragments of DNA. This process continues simultaneously to leading strand synthesis, so there are many small replicating double-stranded DNA fragments on the lagging strand. These are called Okazaki fragments, after the husband and wife duo of Reiji and Tsuneko Okazaki, who discovered them.
Are We Done Yet?
DNA replication ends in bacteria and viruses with circular genomes when the two replication forks meet each other on the opposite side of the chromosome from the origin of replication, in a magnificent game of chicken. The brave replication terminator protein steps in and pauses DNA replication so that they don't crash into each other. However, the mechanism of termination leads to two interlocked circular chromosomes stuck to each other like magician's rings. Topoisomerase enzymes, such as DNA gyrase and topoisomerase IV, which are responsible for nicking DNA, temporarily cut one chromosome so that the other can pass through. There we go, ruining magic tricks for everyone.Animation of Linear Chromosome Shortening
Linear chromosomes have another set of problems because with these guys, we never reach complete replication of the full chromosome. Each replication step shortens the DNA a little bit more. Bacteria and viruses with linear chromosomes get around this problem by either- Initiating replication in the center of the chromosome, or by
- Priming replication with a protein instead of an RNA primer sequence.
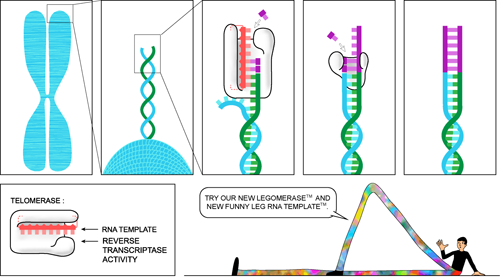
The ends of linear eukaryotic DNA are called telomeres, and though they are regularly shortened every replication cycle, enzymes called telomerases fix the problem of DNA shortening. Telomeres are highly repetitive, and telomerases add short sequences to lengthen the end of DNA so that they do not become too short, and so that important parts of the DNA sequence are not lost. Telomerases are reverse transcriptases, or enzymes (we can tell by the —ase) that convert RNA to DNA. See the Spiderman and Other Examples of Recombinant DNA" section for more.
Telomerases take a short RNA sequence, convert it to double-stranded DNA sequence, and attach it to the end of the DNA. There is a minimum length for telomeres to maintain, and if the DNA sequence is shorter than the minimum length, called the Hayflick limit, the cell dies. Telomerases ensure that the cell can continue to replicate and divide indefinitely. And, overactivity of telomerases is commonly linked to cancer, as cells become immortal, like The Highlander.
Whoops, Mistakes Were Made: Proofreading and Repair
You are probably thinking, "DNA polymerase thinks it is so great because it replicates all the DNA in my body so I can survive and continue to undergo mitosis and meiosis." OK, We are sure most of you are not thinking that, but let's play along to the handful of people that do think that.Actually, DNA polymerase is not that great! It makes mistakes like you or Shmoop—though our mistakes are mostly the products of too many 5-hour energy drinks mixed with deep-fried Twinkies. However, DNA polymerase must fix these mistakes, while we at Shmoop go into a sugar-induced coma.
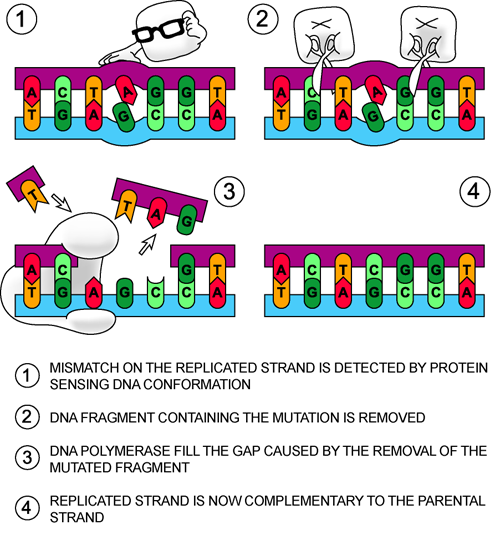
Sometimes, in the haste of DNA replication, a wrong base that does not properly base pair with the template strand is added to the sequence. Therefore, we get weird G-T or A-G base pairs. We all remember that normal pairs are G-C and A-T (Eh? What's that?). These incorrect pairs are not good for us because it means that our genes are mutated, and we become mutants. Few mutations ever lead to awesome things like teleportation or laser vision...or anything out of X-Men. Shucks.
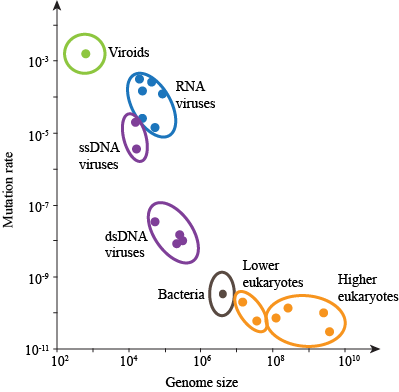
Big eukaryotes like us that replicate slowly and live long don't want much mutation to happen. Why? Any mutation that affects our growth is usually...not good...and will hurt us. Sad face.
However, smaller organisms, like bacteria, viruses, and even flies, have short life cycles on the order of days, and sometimes, hours. They replicate like crazy, where each new generation will have 100s-1000s more individuals than the previous generation. Therefore, mutations in their cases may be more advantageous. Sometimes, they will want to escape pesticides, antibiotics, or drugs designed to treat/kill them, which is why it is so hard to make a drug that will keep us from getting sick. Bacteria and viruses keep mutating and replicating. And, as we have already noticed, there is little error control in their polymerases.
Brain Snack
Here is a neat video of how DNA replication actually works.
Can you spot the DNA polymerases? They are the multicolored spheres, one for the leading strand, and one for the lagging strand. The lagging strand polymerases are the ones coming in and then falling off when the double strand is completed.