ShmoopTube
Where Monty Python meets your 10th grade teacher.
Search Thousands of Shmoop Videos
Molecular Genetics: RNA 245 Views
Share It!
Description:
In this video from our course on molecular genetics learn all about RNA.
Transcript
- 00:00
[ whoosh ]
- 00:01
We speak student!
- 00:03
[ whoosh ]
- 00:04
[ music ]
- 00:06
Molecular Genetics
Full Transcript
- 00:08
RNA
- 00:10
A la Shmoop
- 00:11
[ music continues ]
- 00:13
We are here with Dr. Ruth Tennen
- 00:15
to talk about molecular genetics
- 00:17
here at Shmoop global headquarters
- 00:18
in Mountain View, CA.
- 00:20
And we're gonna cover genetic code and DNA/RNA structure.
- 00:24
All right, so, moving right along,
- 00:26
what is RNA? We've talked about DNA.
- 00:28
What is ribonucleic acid?
- 00:31
Yeah, so, it's very similar to DNA,
- 00:33
but it has that extra oxygen.
- 00:34
So it's also a polymer of nucleotides.
- 00:39
One big difference is, rather than the thymine
- 00:41
as one of the bases, it's uracil. So "T" goes to "U."
- 00:45
- Got it. - But they're basically the same.
- 00:46
Otherwise, their function's very different.
- 00:48
And there's other differences as well. So, for example,
- 00:50
DNA is always double-stranded,
- 00:52
RNA tends to be single-stranded, then fold back on itself.
- 00:55
So they have different functions in the cells,
- 00:57
but they, structurally, are pretty similar.
- 00:58
Got it. Okay.
- 01:00
Then we have subsets of RNA -
- 01:05
mRNA, rRNA, tRNA.
- 01:07
Can you explain to us what are those elements
- 01:10
and how do they function?
- 01:11
[ whooshing ]
- 01:12
What are the different kinds of RNA?
- 01:15
So, kind of a major RNA that we always hear about is mRNA,
- 01:19
which is the intermediate between DNA and protein.
- 01:21
So DNA gets transcribed into mRNA, which gets translated into protein.
- 01:25
So that's kind of like the "famous" RNA.
- 01:27
But there are what's called non-coding RNAs,
- 01:30
that aren't turned into proteins, basically.
- 01:32
So tRNAs are transfer RNAs and those are involved in protein synthesis.
- 01:36
They help with the ribosome.
- 01:39
There are ribosomal RNAs which, again,
- 01:40
they don't get turned into proteins, but they actually
- 01:42
function as RNAs.
- 01:45
But walk us through, like, how does it work?
- 01:48
What's the RNA process? Why do we need these interstitial steps
- 01:51
to make things happen so that everything's okay?
- 01:54
Is it like a human checking mechanism to be sure that genes
- 01:57
are all happy and healthy?
- 01:58
And if they don't, the cell goes away,
- 02:01
so we heal properly?
- 02:03
[ whooshing ]
- 02:04
Why do we need RNA?
- 02:06
One reason is that DNA -- you want it to be super stable.
- 02:09
Right? Because it gets passed down through all the generations
- 02:12
- and we need to keep it the same. - Sure.
- 02:14
By having an intermediate, basically you can say,
- 02:16
"Okay, I wanna turn on this gene.
- 02:18
And I'm gonna make a ton of copies of the messenger RNA."
- 02:20
So you can have a ton of different copies
- 02:21
that ultimately will get degraded when they're not needed anymore.
- 02:24
Whereas you wouldn't wanna chew up the DNA
- 02:26
when it wasn't needed anymore.
- 02:27
So it's kind of a way of...
- 02:29
How do they get degraded?
- 02:29
So this is like taking a DVD and copying it,
- 02:32
and copying it again and again and again
- 02:34
and after a while it just -- like the data isn't as clear?
- 02:38
So each copy, actually, is originally from the DNA.
- 02:40
So each copy's just as good as the previous one.
- 02:42
But, eventually, let's say a cell wanted to turn on a gene
- 02:46
to do a certain thing.
- 02:47
So it said, "Okay, I'm gonna need to metabolize sugar right now
- 02:50
so I gotta turn on my sugar-metabolizing gene."
- 02:52
And then the sugar goes away;
- 02:53
it doesn't need that protein anymore that would do that.
- 02:56
So then there are enzymes that come in
- 02:57
and just chew up and get rid of the RNA. It's extra.
- 02:59
[ woo-woo-woo ]
- 03:01
Got it. So it degrades not in its quality gets less;
- 03:05
it degrades in that its utility is lower.
- 03:07
Exactly. The abundance ends up going down
- 03:09
when it's not needed anymore.
- 03:09
Got it. And our body has stuff that regulates that?
- 03:13
Is that like the...?
- 03:14
You know, I don't know in terms of actual hormones and things,
- 03:17
but at the cellular level, there's like a million enzymes
- 03:19
that are always pucking around and --
- 03:21
Basically, these processes are super important,
- 03:22
so there's tons of layers of regulation
- 03:24
just to make sure everything's pretty tightly constrained.
- 03:26
Got it. 'Cause you've got a lot of replications happening.
- 03:28
I mean, it kind of reminds me of a computer.
- 03:29
It's like if you don't have lots of checks in your database,
- 03:31
- Exactly. - you're gonna print bad data.
- 03:33
[ whoop ]
- 03:34
What is RNA?
- 03:37
What are the different kinds of RNA?
- 03:41
Why do we need RNA?
- 03:45
[ woo-woo-woo ]
Up Next
In this video, we dive beneath the sea to review the kinds of interesting animals that live in the deep blue.
Related Videos
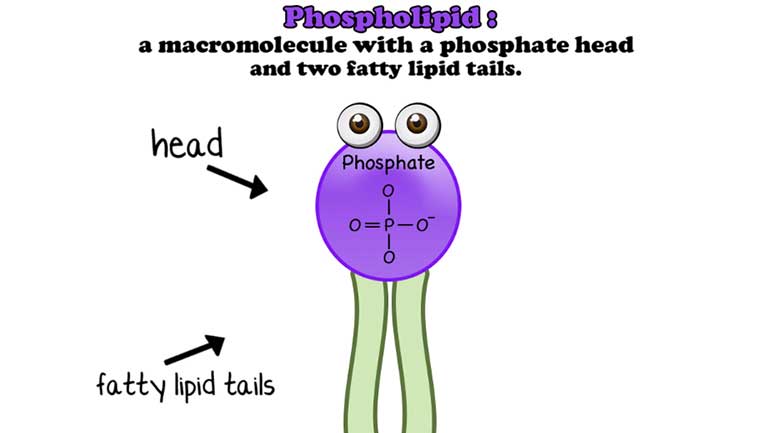
Anything that has a cell (bacteria, listen up!) has phospholipids that keep the cell contained and give it form and shape. Phospholipids protect us...
GMOs. Now that’s a scary word. Or is it? Guess it’s time to ask ourselves: WWMST? ...For those of us who don’t constantly ask ourselves “wh...