Genetic Linkage
Mendel's second law, the law of independent assortment, concerns the random separation of different genes carried on different chromosomes. During meiosis, each chromosome in a homologous pair makes its way into a gamete. Which chromosome from each homologous pair is sorted together into the same gamete is determined by pure chance, so characters on different chromosomes are inherited independently. This is the basis of the law of independent assortment.
But what happens to genes that are on the same chromosome? These genes are physically linked, so they kind of have to go together into the same gamete. They are not inherited independently of one another like genes on different chromosomes, so as well as being physically linked, they are also said to be genetically linked.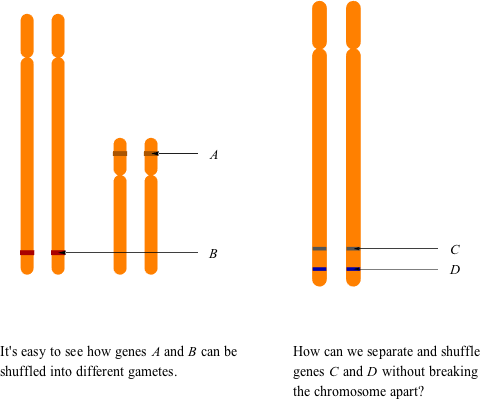 In a way, it's a lot like taking a school trip to the museum. You are part of your class and are "linked" to your classmates. If your class goes to the dinosaur exhibit first, you go with them. Then you go as a group to the watch the 3D show on mammals. At the end of the day, you obviously get on the bus assigned to your class, not the one assigned to the grade above you. That would not be a fun ride—they are not your buddies!
In a way, it's a lot like taking a school trip to the museum. You are part of your class and are "linked" to your classmates. If your class goes to the dinosaur exhibit first, you go with them. Then you go as a group to the watch the 3D show on mammals. At the end of the day, you obviously get on the bus assigned to your class, not the one assigned to the grade above you. That would not be a fun ride—they are not your buddies!
When groups of genes are linked, the alleles for the genes are linked too, and those traits tend to be inherited together. Let's look at a simple real life example.
In the pea plant, the locus for flower color and the locus for pollen shape are on the same chromosome; they are linked. For flower color, purple (symbolized by P) is dominant to red (symbolized by p). For pollen shape, long pollen (symbolized by L) is dominant to round pollen (symbolized by l) (Russell, 1998).
If we cross a true-breeding plant with purple flowers and long pollen (a PPLL genotype that produces only PL gametes) to a true-breeding plant with red flowers and round pollen (a ppll genotype that produces only pl gametes), we get an F1 generation with purple flowers and long pollen (since both purple flower color and long pollen are dominant). Every member of the F1 have a PpLl genotype. Don't believe us? Here's the proof.
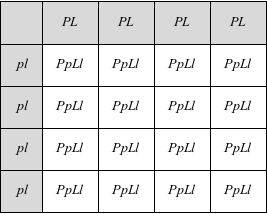
If the genes for flower color and pollen shape are not linked, then the F2 generation will produce our classic 9:3:3:1 phenotypic ratio that we learned about in the Mendelian Genetics section. Just like we did before, let's think of it as a bag of marbles for each locus: a bag for flower color with equal numbers of P and p marbles and a bag for pollen shape with equal numbers of L and l marbles. For each gamete, you blindly draw a marble from each bag. You would expect to see an equal number of gametes with PL, Pl, pL, and pl genotypes. Like this:

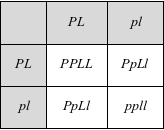
So far, so good, but what happens if they are linked (i.e., on the same chromosome)? In that case, there's a single bag of marbles to deal with. Our bag now holds marbles for both flower color and pollen shape, and each marble has two letters on it. As such, the gametes an F1 produces are only of two kinds: PL and pl. These are the exact same gametes that the parental strains produced because the alleles were linked in each parent. So in the F2 we find only three possible genotypes and two phenotypes:
Okay, so that's the basic theory. But you know how it goes, there's always a catch. As it turns out, linked alleles can become unlinked—they part ways and get into different gametes. If we go back to our school trip comparison, you went everywhere with your class as expected, but then you decided your group was getting kind of boring, so you rode back on a different bus with a different class. No matter what we said earlier, you're not physically linked to your classmates (that would be a bit weird), so you can easily decide to do your own thing.
But how does a gene, which is part of a chromosome, go its own way? During meiosis, the process of gamete formation, homologous chromosomes sometimes swap pieces, an "I'll give you my right arm if you give me yours" kind of thing. Exchanging of identical fragments between homologous chromosomes is called recombination. This process generates new combinations of previously "linked" alleles. The larger the distance between genes on the same chromosome, the more chance there is that recombination will take place between them.
In the image below, D is far away from E and F. We would expect the frequency of recombination of D to be high relative to E and F. We'd also expect E and F to usually travel together because of the short distance between them.

Let's go back to our pea plants and their linked genes for flower color and pollen shape. Picture a heterozygous plant for both loci: one chromosome carries a P allele for flower color and an L allele for pollen shape, while the second chromosome carries a p allele for flower color and an l allele for pollen shape.
Without recombination, this plant produces only two kinds of alleles: 50% PL and 50% pl (just like we discussed before). If self-fertilized, 75% of the next generation (F2) should have purple flowers and elongated pollen (either PPLL or PpLl), and 25% of them should have red flowers and round pollen (ppll).
In reality, because during gamete formation the chromosome pair bearing these genes will sometimes swap pieces, we will also get a small number of plants that combine the different phenotypes in a new way: purple flowers with round pollen and red flowers with elongated pollen. This is because the recombination that took place during meiosis leads to gametes with new allele combinations (either Pl or pL).
Brain Snacks
But what happens to genes that are on the same chromosome? These genes are physically linked, so they kind of have to go together into the same gamete. They are not inherited independently of one another like genes on different chromosomes, so as well as being physically linked, they are also said to be genetically linked.
 In a way, it's a lot like taking a school trip to the museum. You are part of your class and are "linked" to your classmates. If your class goes to the dinosaur exhibit first, you go with them. Then you go as a group to the watch the 3D show on mammals. At the end of the day, you obviously get on the bus assigned to your class, not the one assigned to the grade above you. That would not be a fun ride—they are not your buddies!
In a way, it's a lot like taking a school trip to the museum. You are part of your class and are "linked" to your classmates. If your class goes to the dinosaur exhibit first, you go with them. Then you go as a group to the watch the 3D show on mammals. At the end of the day, you obviously get on the bus assigned to your class, not the one assigned to the grade above you. That would not be a fun ride—they are not your buddies!When groups of genes are linked, the alleles for the genes are linked too, and those traits tend to be inherited together. Let's look at a simple real life example.
In the pea plant, the locus for flower color and the locus for pollen shape are on the same chromosome; they are linked. For flower color, purple (symbolized by P) is dominant to red (symbolized by p). For pollen shape, long pollen (symbolized by L) is dominant to round pollen (symbolized by l) (Russell, 1998).
If we cross a true-breeding plant with purple flowers and long pollen (a PPLL genotype that produces only PL gametes) to a true-breeding plant with red flowers and round pollen (a ppll genotype that produces only pl gametes), we get an F1 generation with purple flowers and long pollen (since both purple flower color and long pollen are dominant). Every member of the F1 have a PpLl genotype. Don't believe us? Here's the proof.

If the genes for flower color and pollen shape are not linked, then the F2 generation will produce our classic 9:3:3:1 phenotypic ratio that we learned about in the Mendelian Genetics section. Just like we did before, let's think of it as a bag of marbles for each locus: a bag for flower color with equal numbers of P and p marbles and a bag for pollen shape with equal numbers of L and l marbles. For each gamete, you blindly draw a marble from each bag. You would expect to see an equal number of gametes with PL, Pl, pL, and pl genotypes. Like this:

So far, so good, but what happens if they are linked (i.e., on the same chromosome)? In that case, there's a single bag of marbles to deal with. Our bag now holds marbles for both flower color and pollen shape, and each marble has two letters on it. As such, the gametes an F1 produces are only of two kinds: PL and pl. These are the exact same gametes that the parental strains produced because the alleles were linked in each parent. So in the F2 we find only three possible genotypes and two phenotypes:

Okay, so that's the basic theory. But you know how it goes, there's always a catch. As it turns out, linked alleles can become unlinked—they part ways and get into different gametes. If we go back to our school trip comparison, you went everywhere with your class as expected, but then you decided your group was getting kind of boring, so you rode back on a different bus with a different class. No matter what we said earlier, you're not physically linked to your classmates (that would be a bit weird), so you can easily decide to do your own thing.
But how does a gene, which is part of a chromosome, go its own way? During meiosis, the process of gamete formation, homologous chromosomes sometimes swap pieces, an "I'll give you my right arm if you give me yours" kind of thing. Exchanging of identical fragments between homologous chromosomes is called recombination. This process generates new combinations of previously "linked" alleles. The larger the distance between genes on the same chromosome, the more chance there is that recombination will take place between them.
In the image below, D is far away from E and F. We would expect the frequency of recombination of D to be high relative to E and F. We'd also expect E and F to usually travel together because of the short distance between them.

Let's go back to our pea plants and their linked genes for flower color and pollen shape. Picture a heterozygous plant for both loci: one chromosome carries a P allele for flower color and an L allele for pollen shape, while the second chromosome carries a p allele for flower color and an l allele for pollen shape.
Without recombination, this plant produces only two kinds of alleles: 50% PL and 50% pl (just like we discussed before). If self-fertilized, 75% of the next generation (F2) should have purple flowers and elongated pollen (either PPLL or PpLl), and 25% of them should have red flowers and round pollen (ppll).
In reality, because during gamete formation the chromosome pair bearing these genes will sometimes swap pieces, we will also get a small number of plants that combine the different phenotypes in a new way: purple flowers with round pollen and red flowers with elongated pollen. This is because the recombination that took place during meiosis leads to gametes with new allele combinations (either Pl or pL).
Brain Snacks
- Did you know that you can use recombination frequency to work out where genes are on a chromosome? By looking at the numbers of recombinant phenotypes compared to the ones you were expecting, you can work out where the genes are in relationship to each other. The unit of measurement for these chromosomal gene maps is called the centiMorgan (cM), named after the geneticist Thomas Hunt Morgan, who worked on linkage using the fruit fly Drosophila melanogaster.
- Fruit flies are very important in genetic research, but they are a bit pongy, thanks to the fact that they live on rotting fruit. For more on how Drosophila is used to study heredity, have a look at this article from Catalyst magazine